New preprint out. We investigated whether we could detect evidence for genetic recombination in #SARSCoV2, and found none to date. We used (slightly tweaked) standard statistical genetics methods, extensive simulations and a validation in #MERSCoV.
1/ https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
1/ https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
Recombination in #SARSCoV2 has been largely overlooked to date. Recombination-mediated rearrangement of variants that arose independently can be of major evolutionary importance. The absence of recombination is also a key assumption behind any phylogenetic inference.
2/
2/
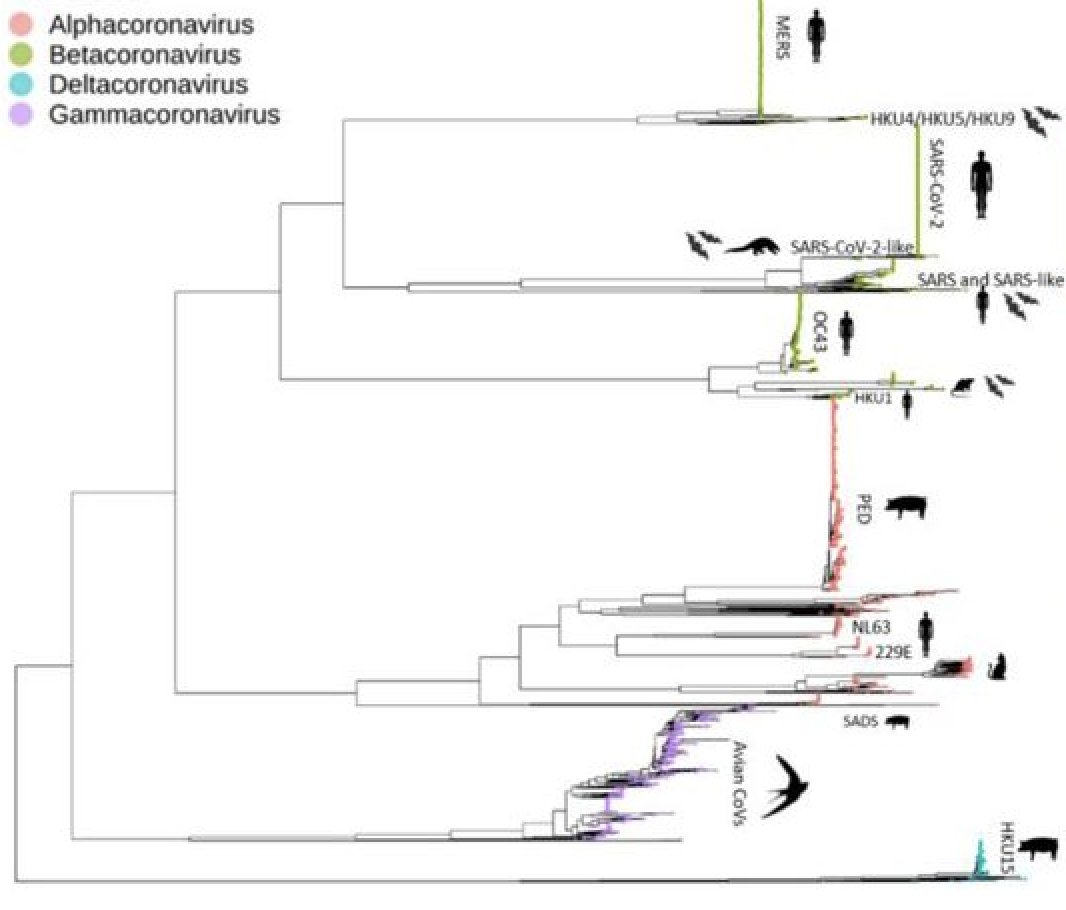
It is often assumed that coronaviruses recombine readily between and within 'species lineages'. We previously found no evidence for recombination between different coronaviruses. #SARSCoV2 in particular doesn't look like a recombinant.
3/
https://www.biorxiv.org/content/10.1101/2020.12.08.415703v1.full
3/
https://www.biorxiv.org/content/10.1101/2020.12.08.415703v1.full
Detecting signals of genetic recombination between lineages belonging to the same species is more challenging. This is particularly the case for #SARSCoV2, which still has low genetic diversity at this stage.
4/ https://www.nature.com/articles/s41467-020-19818-2
4/ https://www.nature.com/articles/s41467-020-19818-2
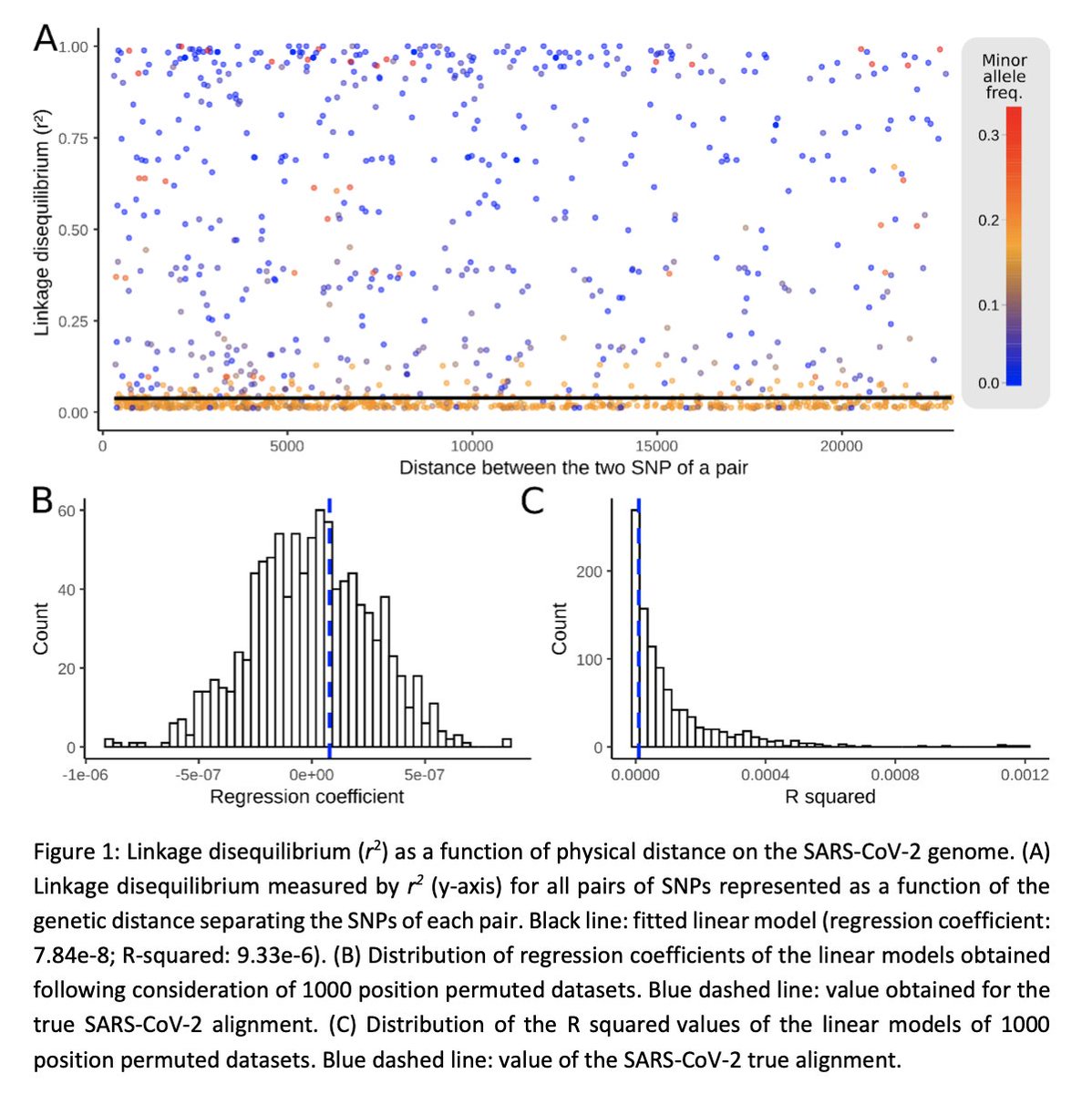
To test for genetic recombination in #SARSCoV2, we relied primarily on the prediction that in the presence of recombination, sites found nearby on the genome are expected to be more strongly correlated than more distant ones.
5/ https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
5/ https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
We found no detectable hallmark of genetic recombination in #SARSCoV2. We run extensive simulations matching the genetic pattern observed in the #SARSCoV2 population today to assess our detection power and validate our method.
6/
https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
6/
https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
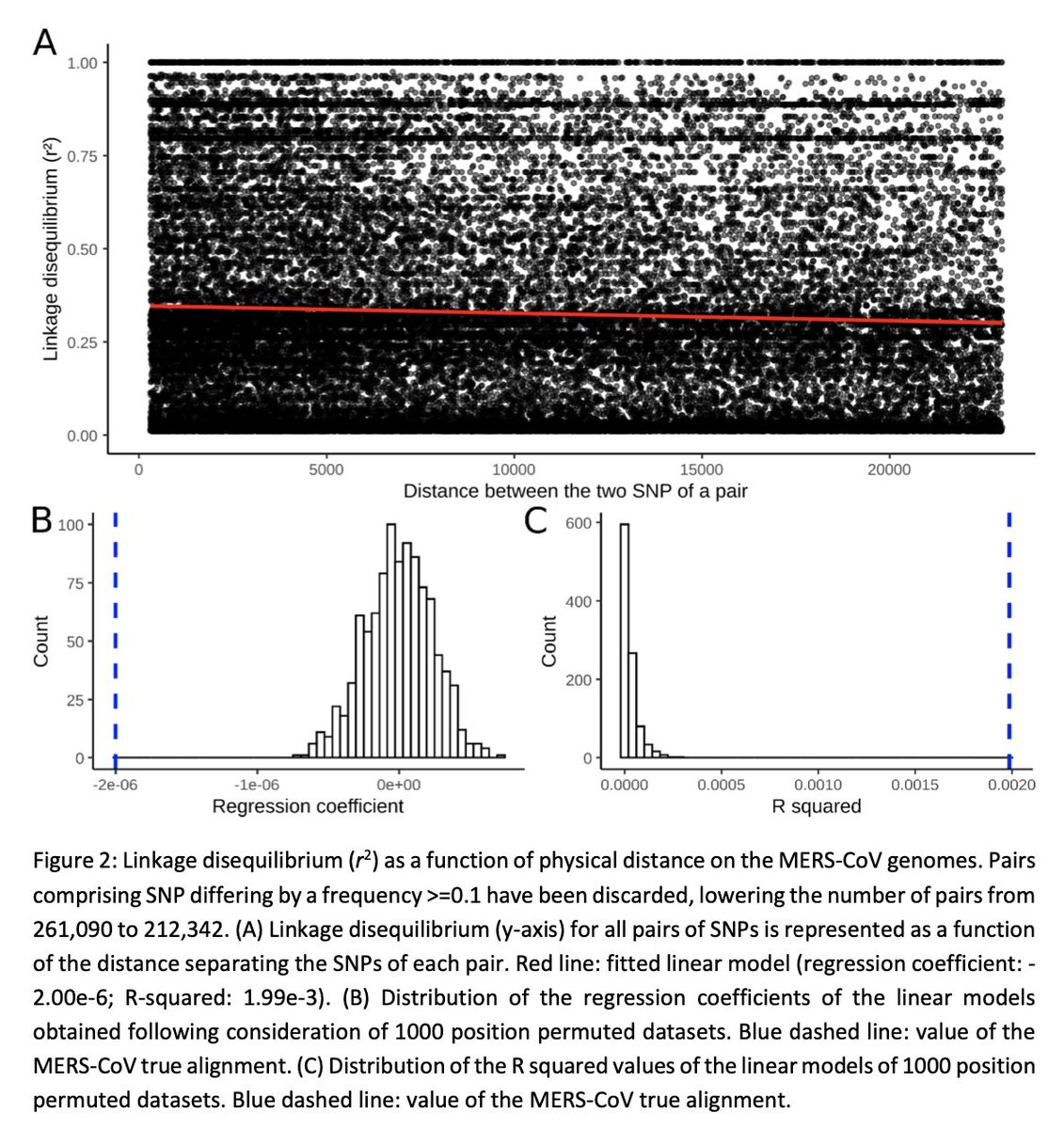
We also validated our method on the related #MERSCoV, another beta-coronaviruses circulating in camelids in the Arabic Peninsula and causing regular 'spillover outbreaks' in humans. We observed a strong signal for genetic recombination in #MERSCoV.
7/
https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
7/
https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
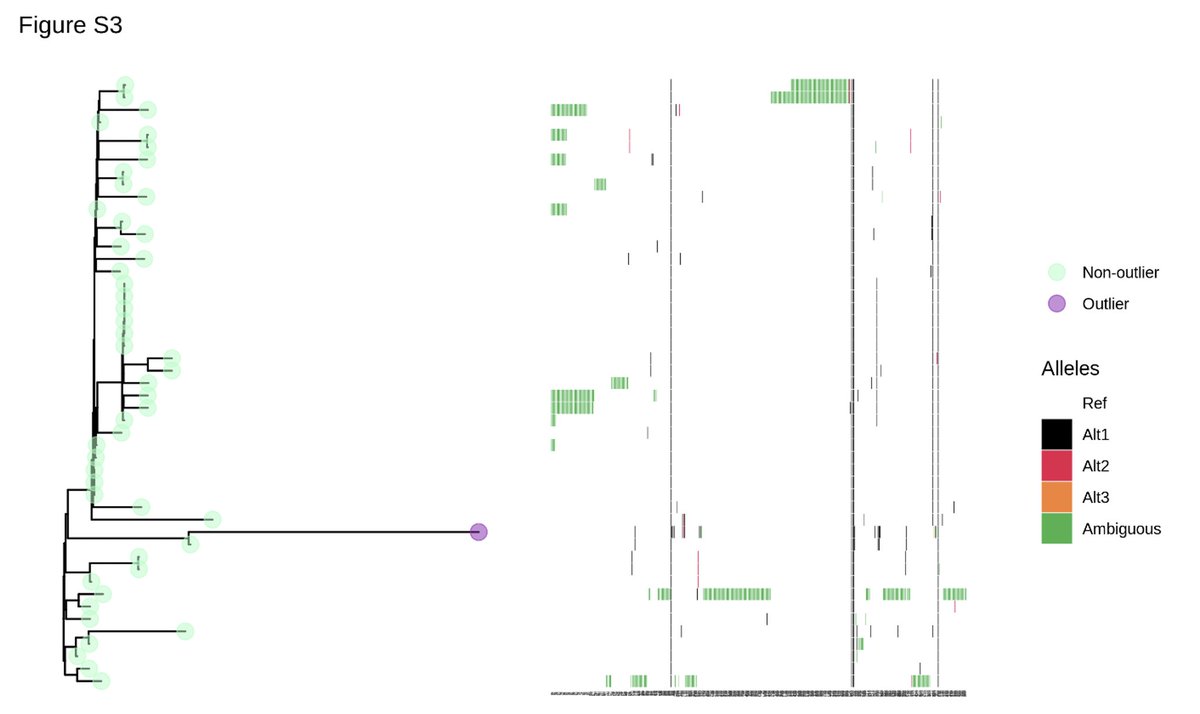
We also analysed in detail #SARSCoV2 genomes on 'long branches' of the phylogeny (i.e. that have accumulated many mutations), as those are plausible candidates for recombination (i.e. chimeric genomes). Again we picked up no signal for recombination.
8/
https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
8/
https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
In summary, we detected no genetic recombination in the #SARSCoV2 population to date. This may partly be due to limited statistical power caused by the still low genetic diversity and the limited potential for mixed infections (≥2 strains) so far.
9/ https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
9/ https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
#SARSCoV2 not being particularly prone to undergo recombination is good news on the vaccine front. Immune-escape mutants generally arise through the accumulation of multiple mutations (antigenic drift), which is a slower process without recombination.
10/ https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
10/ https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
Thank you Damien, @ChrisJ_Owen and @LucyvanDorp. You're a wonderful team, I couldn't dream of working with more fantastic collaborators. We're most grateful to everyone who contributed to generate the genomic data, and in particular to #COGUK.
11/ https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1
11/ https://www.biorxiv.org/content/10.1101/2020.12.15.422866v1

 Read on Twitter
Read on Twitter