Happy to share our latest pre-print about how we identified the barriers that prevent FeS enzymes from functioning within non-native hosts.
If you're a synthetic biologist tired of your cool FeS enzymes not working in model organisms, this is for you!
(1/9) https://www.biorxiv.org/content/10.1101/2021.02.02.429153v1
If you're a synthetic biologist tired of your cool FeS enzymes not working in model organisms, this is for you!
(1/9) https://www.biorxiv.org/content/10.1101/2021.02.02.429153v1
A common issue in #synbio & metabolic eng. is that many enzymes cannot function outside of their native host.
The obstacles that prevent enzymatic activity are rarely studied and frustrate the development of microbial cell factories. This is the case for many FeS enzymes /2
The obstacles that prevent enzymatic activity are rarely studied and frustrate the development of microbial cell factories. This is the case for many FeS enzymes /2
FeS enzymes are highly versatile & play irreplaceable roles in the biosynthesis of natural products, including antibiotic and #antiviral compounds.
So we decided to find out why these enzymes are often inactive in model bacteria such as E. coli and find ways to activate them /3
So we decided to find out why these enzymes are often inactive in model bacteria such as E. coli and find ways to activate them /3
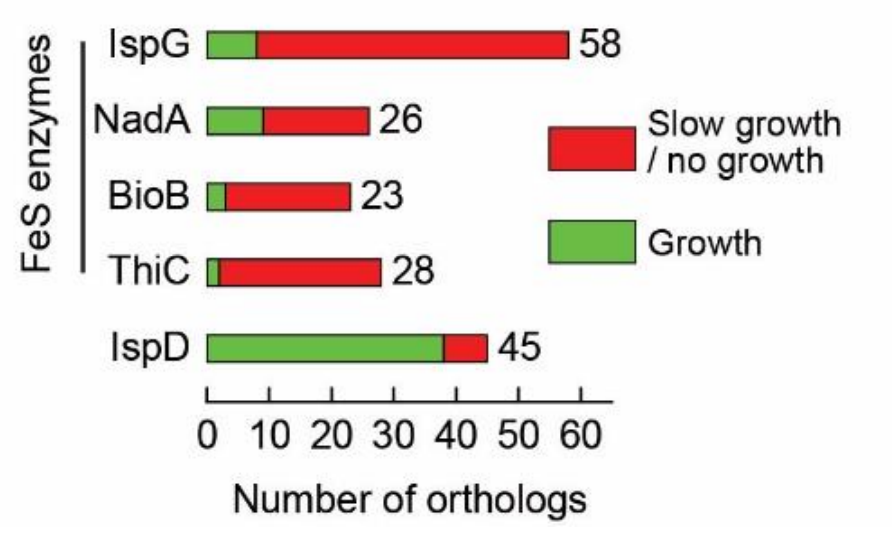
First we develop a simple complementation assay to experimentally test the functionality of orthologs of conditionally-essential FeS enzymes in E. coli.
We show that MOST FeS enzymes from other organisms tested don't work! /4
We show that MOST FeS enzymes from other organisms tested don't work! /4
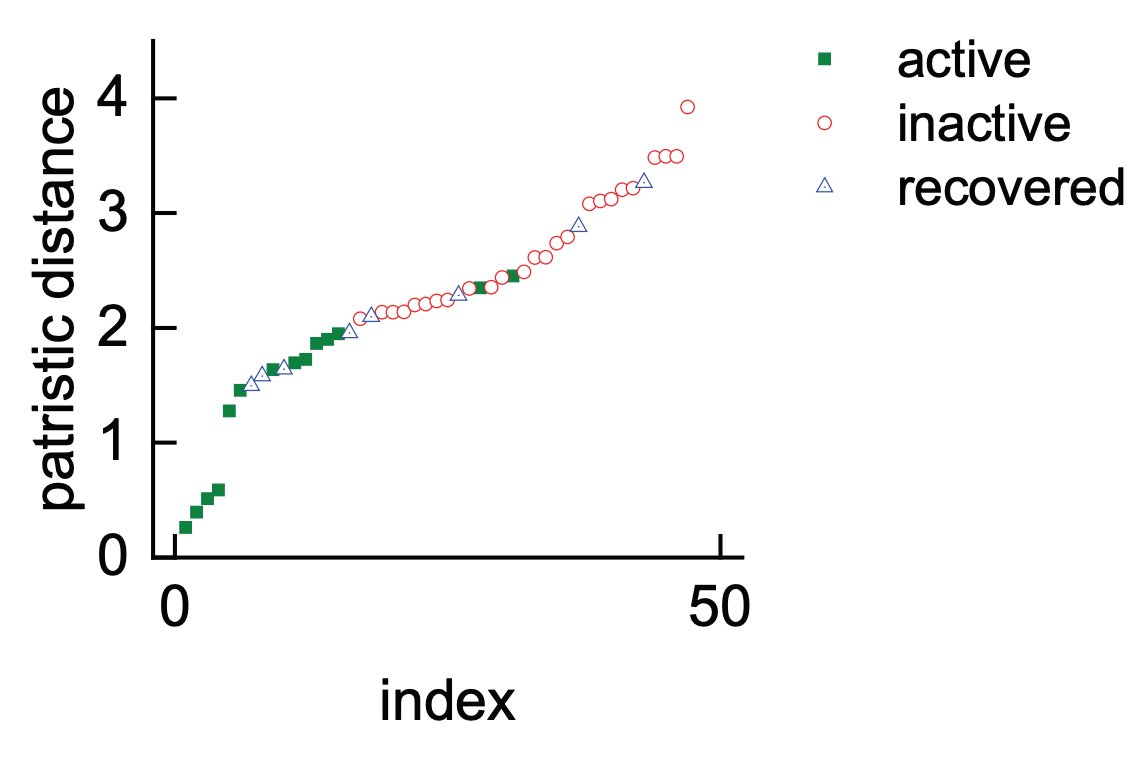
Using FeS enzyme orthologs representative of all of prokaryotic diversity we reveal a clear correlation between the phylogeny of an FeS enzyme and the likelihood of its functioning within a non-native host.
Below for example the patristic distance and NadA activity in E. coli
/5
Below for example the patristic distance and NadA activity in E. coli
/5
We distinguish 2 independent barriers to FeS enzyme activity. The lack of:
- Specific proteins involved in FeS cluster maturation
- Compatible electron transfer protein partners involved in FeS catalysis
Co-expressing such proteins in the host activates foreign FeS enzymes! /6
- Specific proteins involved in FeS cluster maturation
- Compatible electron transfer protein partners involved in FeS catalysis
Co-expressing such proteins in the host activates foreign FeS enzymes! /6
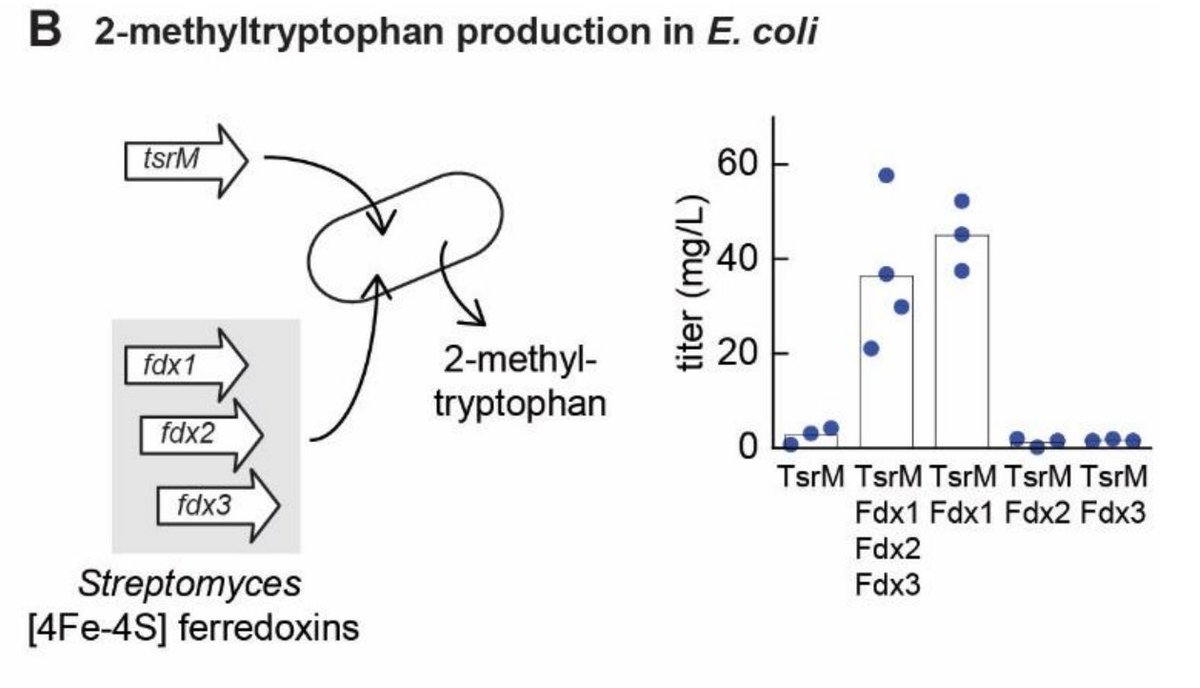
Finally we discover the key cofactors required for in vivo activity of enzyme TsrM, an intensely-studied model enzyme representing the radical SAM family. /7
Our results clarify how phylogenetic distance and electron transfer biochemistry each separately impact functional heterologous expression and provide insight into how these barriers can be overcome for successful microbial engineering involving FeS enzymes. /8
This project was developed during my PhD with Greg Bokinsky @tudelft, and this manuscript is the product of a big interdisplinary collaboration that includes @FredBarras group, Sandrine Ollagnier de Choudens, Beatrice Py, @SGribaldo & more! 9/9

 Read on Twitter
Read on Twitter