An updated version of our report, investigating an association between SARS-COV-2 viral load and mutations associated with the new U.K. variant is linked here.
https://www.coronavirus-fraser-group.org/files/files/updated_report_to_nervtag_oxford_20201222.pdf
https://www.coronavirus-fraser-group.org/files/files/updated_report_to_nervtag_oxford_20201222.pdf
This report has been sent to NERVTAG along with and earlier version (linked in the above document). This was one several pieces of evidence reviewed at that meeting.
https://khub.net/documents/135939561/338928724/SARS-CoV-2+variant+under+investigation%2C+meeting+minutes.pdf/962e866b-161f-2fd5-1030-32b6ab467896?t=1608470511452
https://khub.net/documents/135939561/338928724/SARS-CoV-2+variant+under+investigation%2C+meeting+minutes.pdf/962e866b-161f-2fd5-1030-32b6ab467896?t=1608470511452
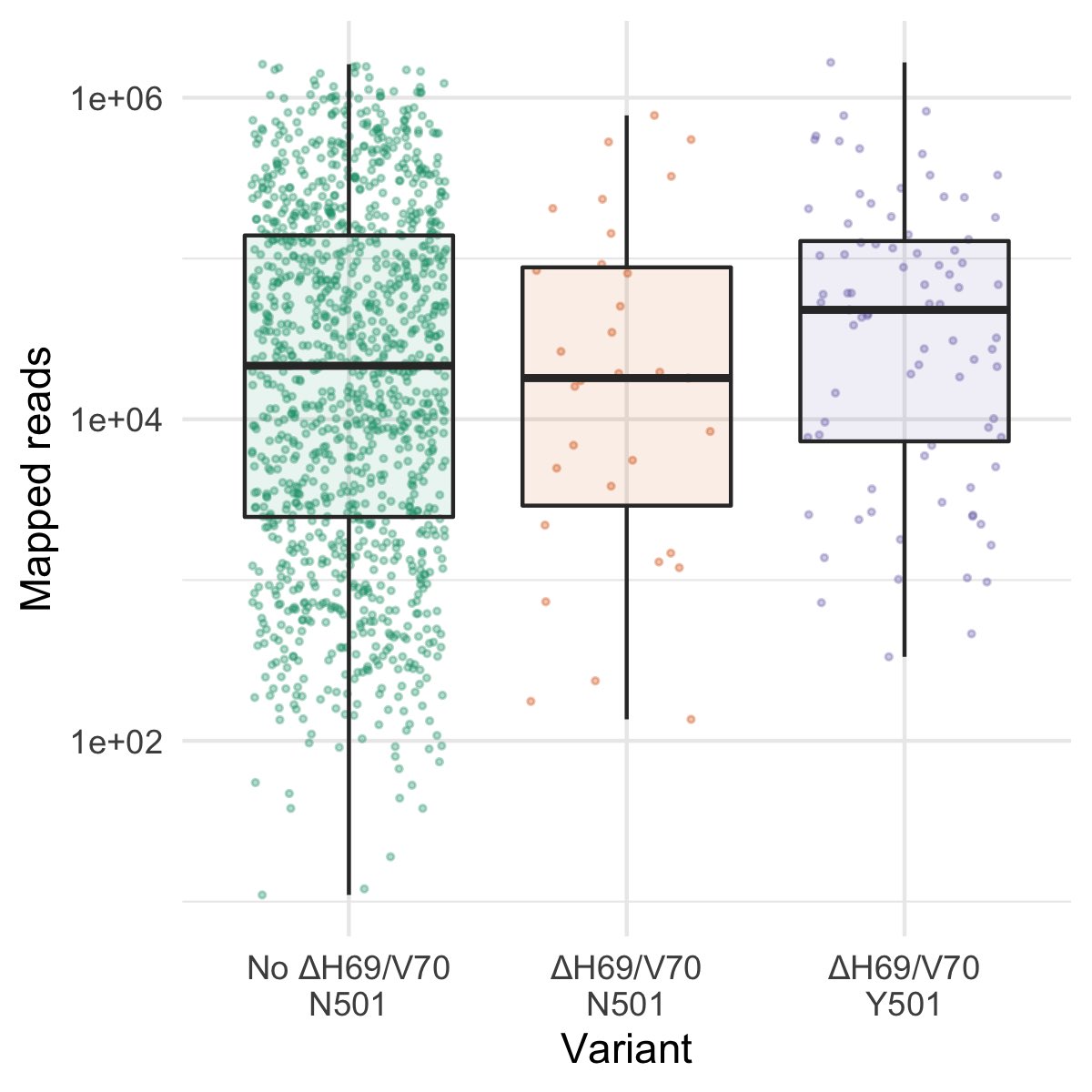
A new variant of SARS-CoV-2 has emerged which is increasing in frequency, primarily in the South East of England (lineage B.1.1.7; VUI-202012/01). One hypothesis is infection with the new variant results in higher viral loads, which in turn may make the virus more transmissible.
We found higher (sequence derived) viral loads in samples from individuals infected with the new variant. Median inferred viral loads were three-fold higher in individuals with the new variant
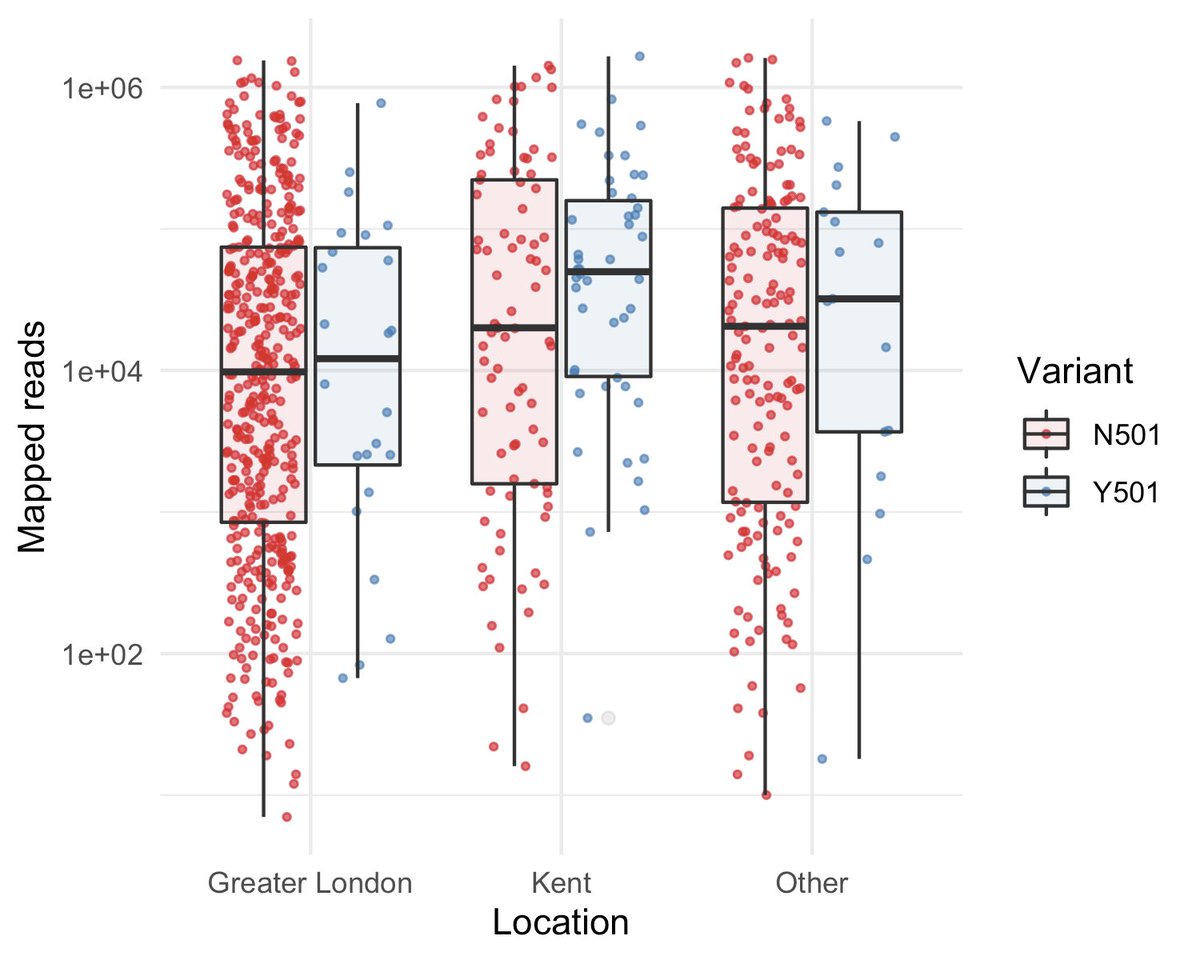
Most of the new variants were sampled in Kent and Greater London. We observed higher viral loads in Kent compared to Greater London for both the new variant and other circulating lineages.
Outside Greater London, the variant has higher viral loads. Within Greater London, the new variant does not have significantly higher viral loads compared to other circulating lineages.
High variant VLs outside Greater London could be due to demographic effects, such as a faster variant growth rate compared to other lineages or concentration in particular age-groups.
Our analysis does not exclude a causal link between infection with the new variant and higher viral loads.
This rapid analysis was possible using a a high throughput sequencing method we developed in Oxford that both sequences whole genomes, and estimates viral load in a single test. It works well on other virus to https://jcm.asm.org/content/58/10/e00382-20
This remains a provisional report, and not a complete scientific study. Under normal circumstances more genomes and metadata would be obtained and included before making this report public, and will be added before pre-printing and submitting for peer review.
Further work is needed to investigate any potential causal link between infection with this new variant and higher viral loads, and whether this results in higher transmissibility, severity of infection, or affects relative rates of symptomatic and asymptomatic infection
Thanks to @katrina_lythgoe , @tanyagolubchik , @ChristoPhraser , @mdhall272, @LucaFerrettiEvo. We’re proud to be a part of @CovidGenomicsUK who provide the funds, infrastructure and support for this work.
Also linked here https://www.coronavirus-fraser-group.org/sequencing

 Read on Twitter
Read on Twitter