I've got a new paper about using the computer to find drugs that might stop coronavirus entering cells in the lab. I should stress that that's very, *very* far removed from finding drugs that treat disease, but we use some cool concepts on the way.
Link: https://www.sciencedirect.com/science/article/pii/S2001037020305213
Link: https://www.sciencedirect.com/science/article/pii/S2001037020305213
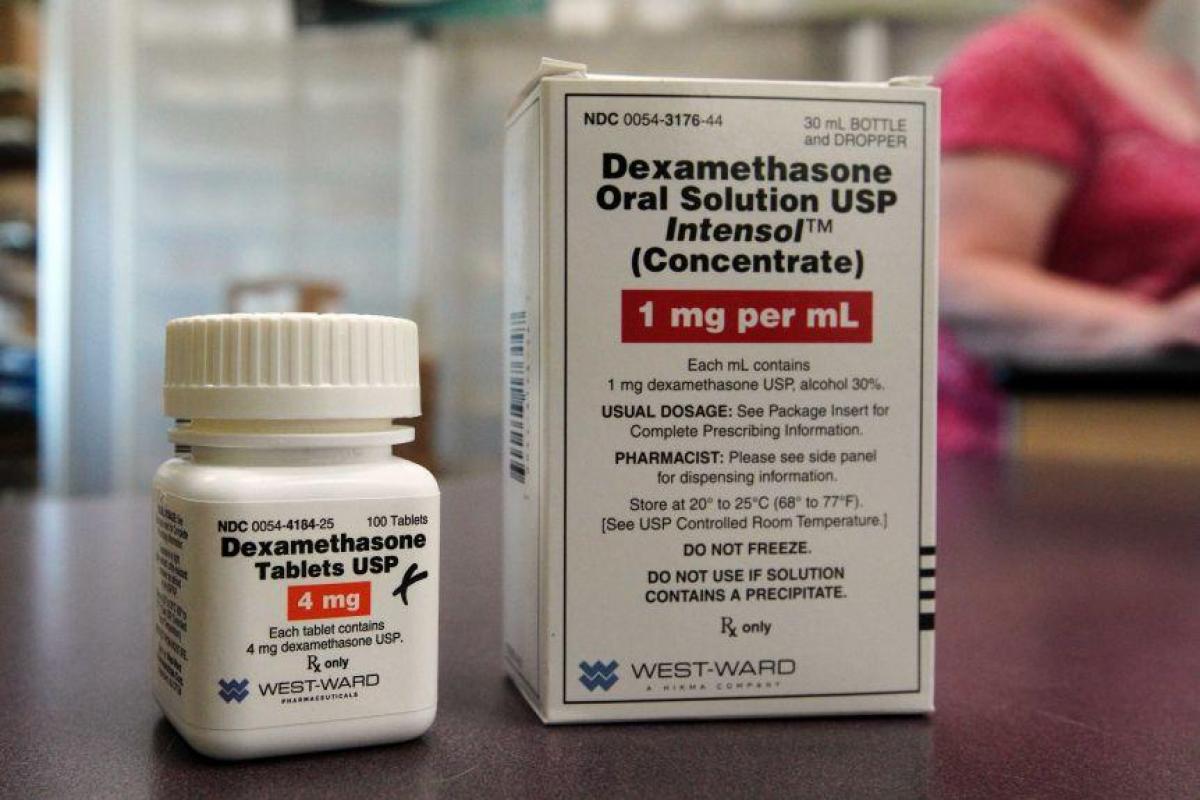
As we know from the RECOVERY trial, we can use dexamethasone, a steroid already in clinical use, to treat selected COVID-19 patients. As it was already in clinical use, we know its safety profile, and the preclinical development was taken care of ages ago.
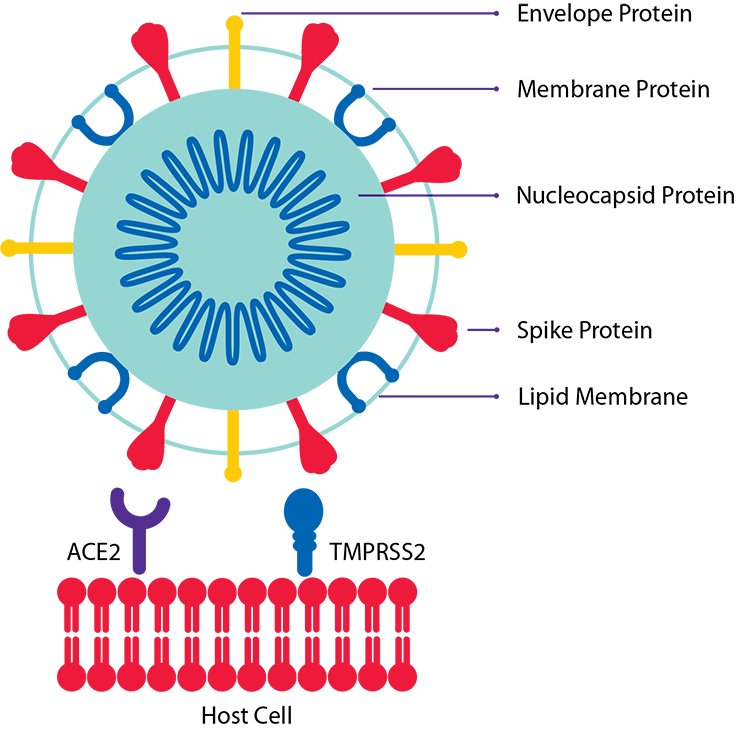
We can also explore more precise treatments now that we know a bit more about the disease. We know, for example, that SARS-CoV-2 infects cells by engaging its spike protein with a molecule called ACE2 that sits in the membrane of a target cell, then entering the cell.
The spike protein has to be just the right shape to engage with ACE2, so a sneaky thing the virus does is use an enzyme on the target cell called TMPRSS2 to whittle the spike. If you block the activity of TMPRSS2, it's harder for the virus to enter cells. https://www.sciencedirect.com/science/article/pii/S0092867420302294
We don't actually know what TMPRSS2 is for. I mean, we know what it does — it cuts up proteins — but we don't know why mammals need to cut proteins in the way TMPRSS2 does. If you genetically engineer mice so they have no TMPRSS2, they seem healthy. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1347042/
So I wondered: are there any drugs that are already in clinical use, whose safety profiles we already know, that might cause our cells to produce less TMPRSS2 so they might be more difficult to infect in the lab? A connectivity map might be a good place to look.
Connectivity maps were developed at MIT with the aim of cataloguing how each drug in a huge library of drugs affected the gene expression profiles of cells. Gene expression profiles give you a rough idea of how much of each protein a cell wants to make.
Recording the behaviour of cells with each drug at each concentration would have taken ages. The clever thing the MIT researchers did, though, was measure only a small portion of the gene expression profile and use a computer model to figure out the rest. https://www.sciencedirect.com/science/article/pii/S0092867417313090
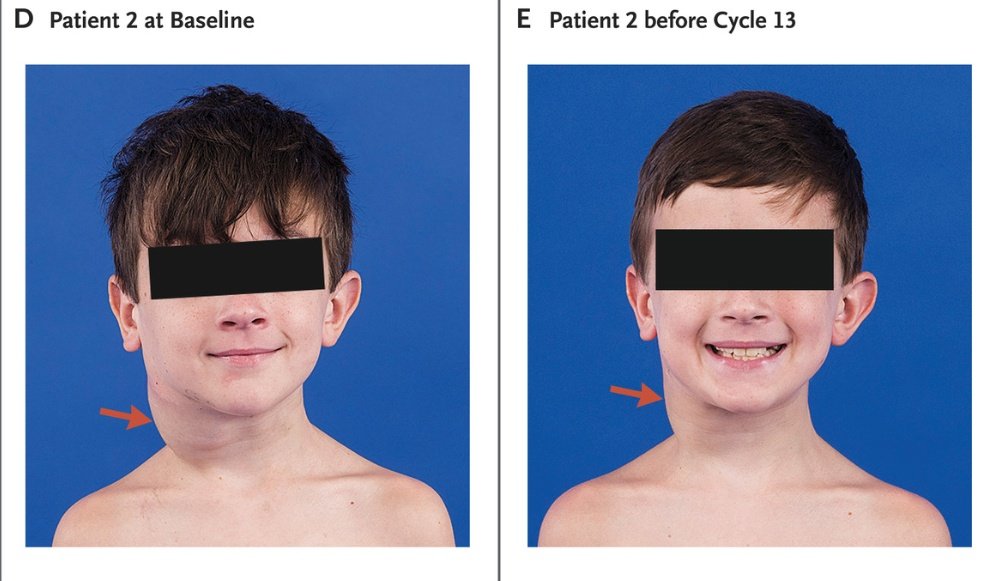
Since they did the hard work, we can search through their database to see if any of the drugs in the library might reduce TMPRSS2 expression. As it turns out, a few appeared to, including a drug from a class known as MEK inhibitors, sometimes used to treat neurofibromatosis.
Does the drug work to stop the virus getting into cells? Does it even have an effect on TMPRSS2 expression? Without doing the experiments we have no clue. Since we're already sitting at the computer, though, we can find out if anyone has seen anything that might suggest it works.
As it happens, in 2014 a group of virologists tested MEK inhibitors to see if they would work to block MERS-CoV cell entry, and they seemed to. MERS-CoV is another coronavirus that uses TMPRSS2 to get its spike into the right shape to infect cells. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4335870/
Those researchers didn't implicate TMPRSS2, but they did figure that early virus-cell interactions might be at play, which fits with our hypothesis.
If MEK inhibitors do actually reduce TMPRSS2 expression, how might they do it? Again, I don't know, but I can hazard a guess.
If MEK inhibitors do actually reduce TMPRSS2 expression, how might they do it? Again, I don't know, but I can hazard a guess.
TMPRSS2 expression is regulated by the androgen receptor, which itself is regulated by a factor called CREB1. MEK inhibitors might block activation of CREB1. It *might* also be the case that the role of the androgen receptor influences how men get more severe Covid than women.
Again, none of this computer work means that we should forgo proper testing in the lab and subsequent clinical trials if the tests are successful. However, if you're looking for a starting point for candidate drugs, there are worse places to look than connectivity maps.
As ever, big thanks to @NIHRNewcBRC, @MedResFdn_AMR and @shieldamr for allowing me to pretend I know any science, and thanks to the Simpson/ @RostronTony teams for allowing me to keep rambling about things I've seen on the computer.

 Read on Twitter
Read on Twitter