I'm convinced ddPCR is more sensitive wrt environmental samples than qPCR. Many assays that we use on the digital platform were designed qPCR but could ddPCR be more sensitive (in WW) if the assays were redesigned according to the instrument's 'tighter' specs?
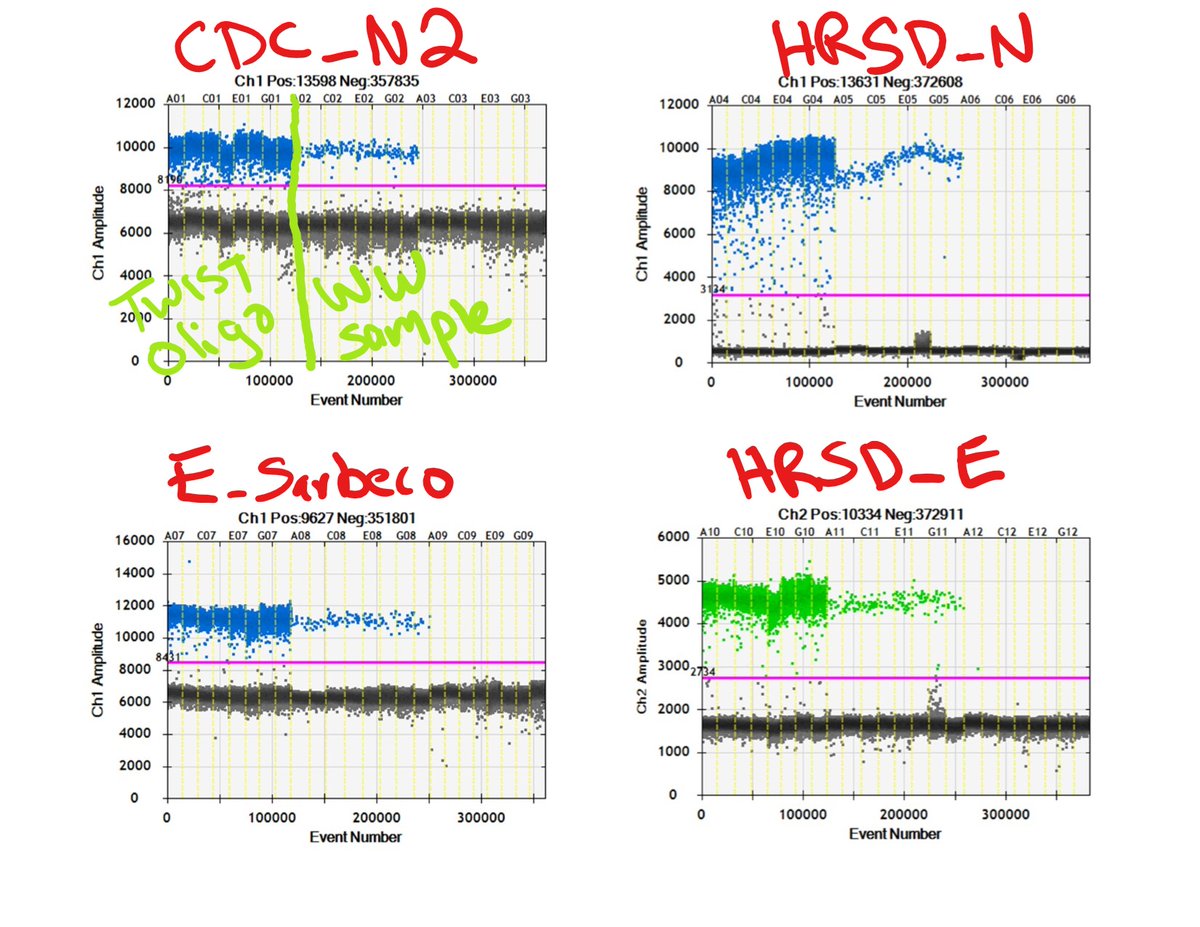
So we designed new E- and N-gene SARS-CoV-2 assays for digital PCR. @PalindromeGirl6 @allycat0089 @novella757 just ran the above annealing temp thermal gradients. Next week we’ll finish LODs and compare WW concentrations.
Takeaways from just the thermal gradients:
-redesigned N gene assay has a way bigger gap between pos and neg droplets
-no improvements on E gene assays but the E Sarbeco assay is showing bleed through to the other channel. This isn't happening on redesign
-redesigned N gene assay has a way bigger gap between pos and neg droplets
-no improvements on E gene assays but the E Sarbeco assay is showing bleed through to the other channel. This isn't happening on redesign
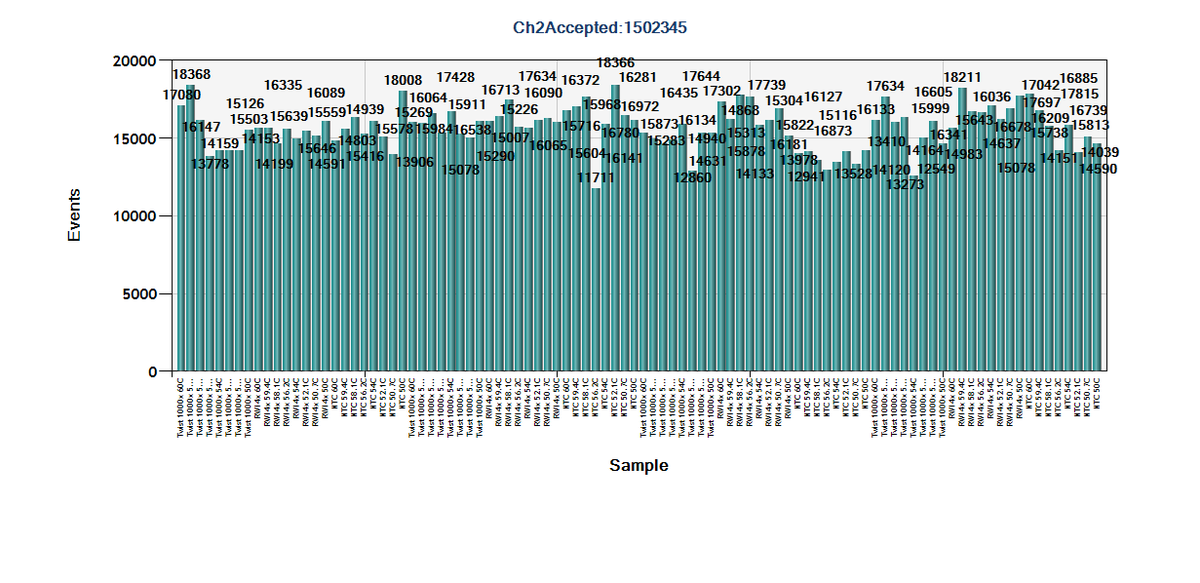
Found their droplet counts (using one step kit) for the plate. Total droplets down but don't see a difference between the twist standard and the wastewater
yes, they run a lot of negatives

 Read on Twitter
Read on Twitter