1/ Was the pandemic caused by SARS2 the result of vaccine research gone wrong? The following thread is the product of a DRASTIC* investigation: the possibility that SARS2 and RaTG13 might be two different vaccine strategies and that SARS2 leaked from a lab during its development.
2/ Which are typical features of a live attenuated vaccine (LAV)? LAV should infect a host with lower virulence and replication capability than the WT, trigger a pronounced adaptive immune response but be cleared quickly without latent infection and it must not revert to its WT.
3/ “Circuit breakers” are required in LAV and the more are integrated into the genome, the safer is the LAV. The resulting fitness attenuation must however retain immunogenicity and protection efficacy against WT challenges.
4/A central mechanism in mammals that controls both innate and adaptive immune responses is mediated by interferons (IFNs). LAV should therefore induce IFN, but rapid clearance after IFN induction would be desirable.
5/ SARS2 seems to have some signatures in its genome which indicate protection strategies, such as IFN-hypersensitivity when compared to the first SARS.
https://www.biorxiv.org/content/10.1101/2020.03.07.982264v1 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7188648/
https://www.biorxiv.org/content/10.1101/2020.03.07.982264v1 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7188648/
6/ Several viral proteins involved in IFN signalling in SARS2 seem to be affected by attenuation, such as NSP3 (less effective deubiquitinating domain due to 24 AA insertion upstream), ORF3b (premature stop codon)
7/ and truncation/changes in ORF6. https://www.biorxiv.org/content/10.1101/2020.03.07.982264v1
8/ Bat immortalized kidney cell line RaK4324 expressing hACE could provide the environment for hACE2 adaptation and avoid at the same time IFN response.
9/ Moreover, the QTQTN motive proximal to the FCS is beneficial for virus entry in presence of Cathepsin, which is naturally produced by kidney cells.
https://s3-eu-west-1.amazonaws.com/itempdf74155353254prod/12213125/Targeting_TMPRSS2_and_Cathepsin_B_L_Together_May_Be_Synergistic_Against_SARS-CoV-2_Infection_v1.pdf
https://s3-eu-west-1.amazonaws.com/itempdf74155353254prod/12213125/Targeting_TMPRSS2_and_Cathepsin_B_L_Together_May_Be_Synergistic_Against_SARS-CoV-2_Infection_v1.pdf
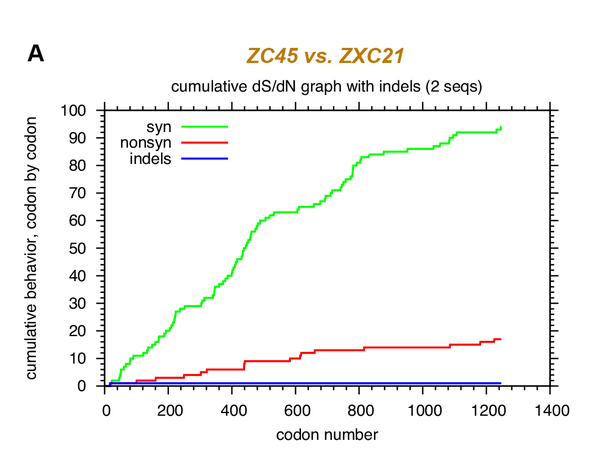
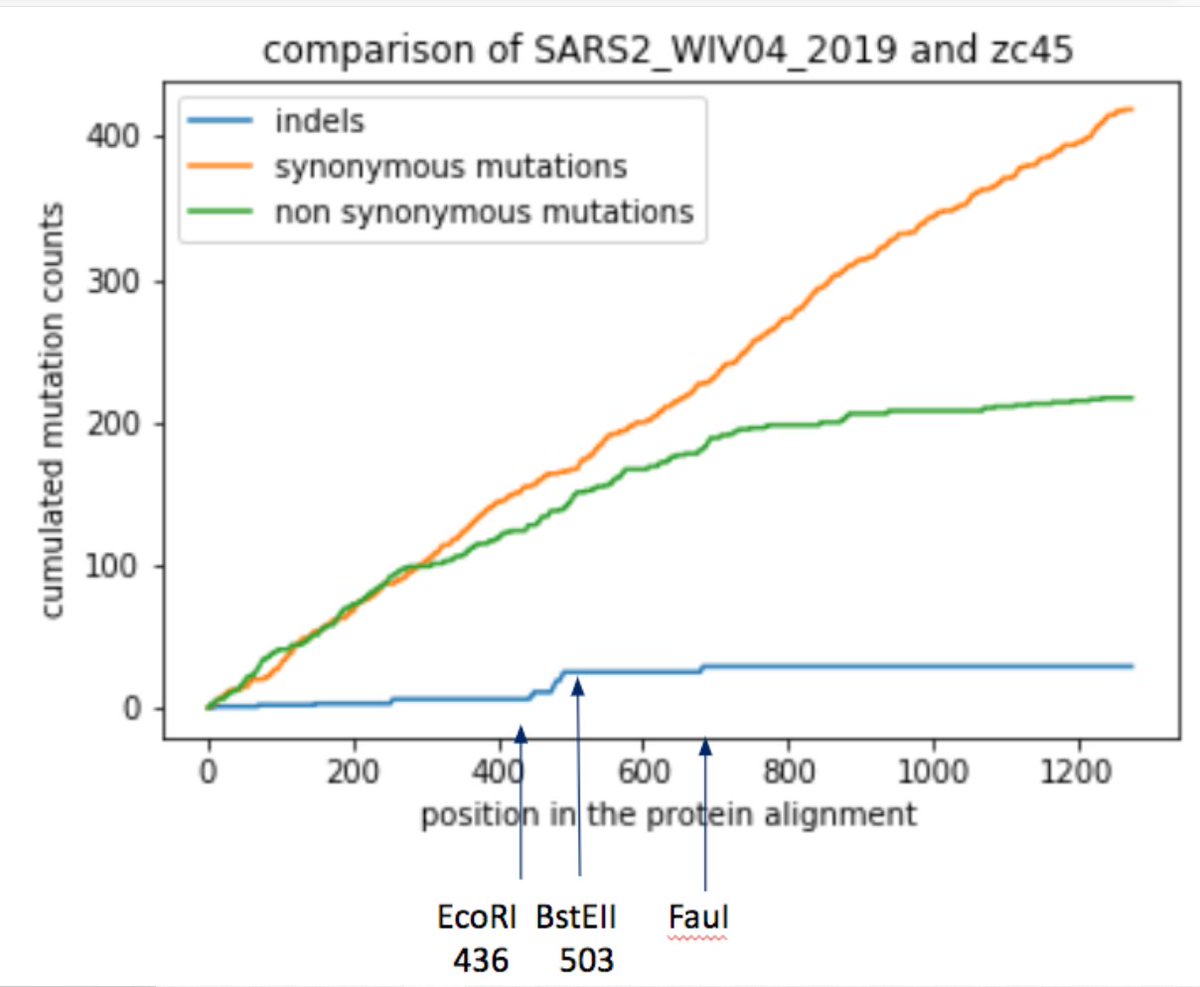
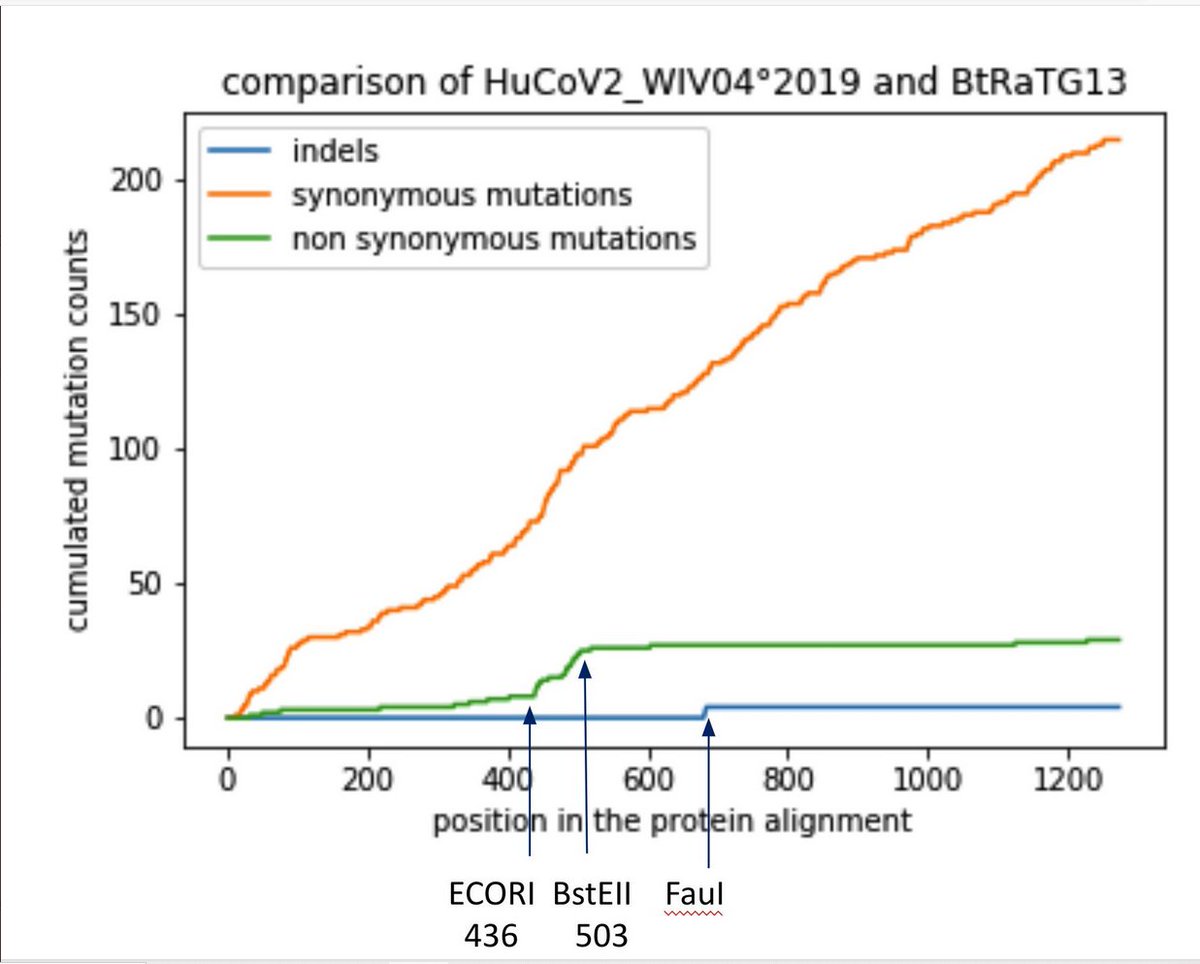
10/ Synonymous mutations were used in the past as strategy for attenuating viruses through codon deoptimization. By comparing RaTG13 with SARS2, an accumulation of synonymous mutations in spike around the FCS is clearly observed.
https://www.researchgate.net/publication/6816952_Reduction_of_the_Rate_of_Poliovirus_Protein_Synthesis_through_Large-Scale_Codon_Deoptimization_Causes_Attenuation_of_Viral_Virulence_by_Lowering_Specific_Infectivity
https://jvi.asm.org/content/89/7/3523
https://www.researchgate.net/publication/6816952_Reduction_of_the_Rate_of_Poliovirus_Protein_Synthesis_through_Large-Scale_Codon_Deoptimization_Causes_Attenuation_of_Viral_Virulence_by_Lowering_Specific_Infectivity
https://jvi.asm.org/content/89/7/3523
11/ Interestingly, the № of CpG dinucleotides of SARS2 is significantly lower than in SARS or MERS and may point as well towards attenuation.
12/ Only in certain regions (such as FCS) there is an accumulation of CpG that could direct an immune response towards S in an otherwise attenuated virus.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3108434/ https://link.springer.com/article/10.1007/s00011-020-01377-3
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3108434/ https://link.springer.com/article/10.1007/s00011-020-01377-3
13/ The use of recombination resistant strains is another strategy for LAV development and it can be achieved by manipulating the viral transcription regulatory networks (TRNs).
14/ In 2018 Baric published a paper where the typical TRS transcription leader and body sequences of SARS were substituted with a novel sequence (UGGUCGC) to reduce recombination in animal models as a live vaccine candidate. https://www.nature.com/articles/s42003-018-0175-7#Sec7
15/ This TRS leader sequence starts at nucleotide 1465 in SARS2 and could result in a novel viral RNA transcript that deletes part of the nsp2 protein in the ORF 1ab polyprotein. https://heraclitusoncovid19.com/origin-of-covid-19/
16/ The same TRS is also found at nucleotide 1446 in RaTG13 but in no other CoVs. Baric refers in his paper to the cave where RaTG13 was collected and RaTG13 was fully sequenced in 2018 but its special TRS is not mentioned. https://www.nature.com/articles/s42003-018-0175-7#Sec7
17/ LAV should also not easily mutate and SARS2 seems to be quite resistant to mutations. This feature might have been achieved by selecting strains with RNA-dependent RNA polymerase (RdRp) with improved fidelity via experiments with mutagenic drugs such as ribavirin.
18/ https://www.nature.com/articles/d41586-020-02544-6 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC165868/
19/ Moreover, ORF10 is coded by a unique sequence in SARS2 and it is not essential. It is immunogenic and it might contribute in hijacking the host’s ubiquitination machinery but it is truncated and probably not expressed, representing another possible attenuation strategy.
20/ https://www.biorxiv.org/content/10.1101/2020.08.29.257360v1
https://www.sciencedirect.com/science/article/pii/S0092867420304062 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2939720/
https://www.sciencedirect.com/science/article/pii/S0092867420304062 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2939720/
21/ How to explain the presence of the FCS? This site (new for Sarbecovirus) could have been inserted to confer immunity not only to SARS-like CoVs but also to CoVs with FCS, such as MERS.
22/ The FCS in SARS2 has highly CpG-rich insertion (CGG-CGG) which is extremely rare as double instance in CoVs and deoptimizes the codon for replication
https://www.researchgate.net/publication/340924249_Is_considering_a_genetic-manipulation_origin_for_SARS-CoV-2_a_conspiracy_theory_that_must_be_censored
https://www.researchgate.net/publication/340924249_Is_considering_a_genetic-manipulation_origin_for_SARS-CoV-2_a_conspiracy_theory_that_must_be_censored
23/ To avoid activation of the FCS, a shield of O-linked glycans was possibly planned, by inserting a leading proline in the sequence, but it failed to be synthetized, probably because of a N-linked glycan site just upstream of the S1-S2, blocking the required glycosyltransferase
24/ https://www.nature.com/articles/s41591-020-0820-9
https://www.cell.com/iscience/pdf/S2589-0042(20)30397-7.pdf
https://www.cell.com/iscience/pdf/S2589-0042(20)30397-7.pdf
25/ If the shield would have been present, not only the FCS would have been blocked but also the activity of Cathepsin and TMPRS2 would have been hampered.
26/ O-glycosylated S1-S2 junction would not be a superantigen and it won’t bind to NPR1, the “door” for the central nervous system.
https://www.pnas.org/content/early/2020/09/25/2010722117 https://www.biorxiv.org/content/10.1101/2020.06.07.137802v3
https://www.pnas.org/content/early/2020/09/25/2010722117 https://www.biorxiv.org/content/10.1101/2020.06.07.137802v3
27/ Testing of stability and effect of FCS in the virus would have required cell lines not prone to its deletion, such as HEK293T, Human Airway Epithelia (HAE) cells or Organ-on-a-chip models.
https://pubmed.ncbi.nlm.nih.gov/30849247/
https://www.biorxiv.org/content/10.1101/2020.09.30.318311v1.full.pdf
https://pubmed.ncbi.nlm.nih.gov/30849247/
https://www.biorxiv.org/content/10.1101/2020.09.30.318311v1.full.pdf
28/ It is not possible to exclude that the original sequence coding for the FCS was designed to be inactive, but it mutated during human to human transmission.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7379507/pdf/fgene-11-00783.pdf
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7379507/pdf/fgene-11-00783.pdf
29/ Moreover, the FCS of CoV2 matches exactly to that of a Heparan Sulphate binding motif—XBBXBX.
https://www.sciencedirect.com/science/article/pii/S0092867420312307
This particular feature confers to SARS2 the ability to efficiently bind to proteins. https://twitter.com/BillyBostickson/status/1276420129346056198
https://www.sciencedirect.com/science/article/pii/S0092867420312307
This particular feature confers to SARS2 the ability to efficiently bind to proteins. https://twitter.com/BillyBostickson/status/1276420129346056198
30/ Not only ACE2 and TMPRSS2 expression, but also Heparan Sulphate binding could explain the striking gradient of SARS-CoV-2 infection in proximal (high) versus distal (low) pulmonary epithelial cultures.
https://www.cell.com/cell/pdf/S0092-8674%2820%2930675-9.pdf https://www.nature.com/articles/s41591-020-0868-6
https://www.cell.com/cell/pdf/S0092-8674%2820%2930675-9.pdf https://www.nature.com/articles/s41591-020-0868-6
31/ In the hypothesis of SARS2 being developed as intranasal or self- spreading vaccine, this ability could reduce the viral load in the lung, where the FCS is more prone to be activated. https://www.sciencedirect.com/science/article/pii/S1097276520302641
32/ but it could also result in adverse reactions in some individuals, in particular those with an impaired antiviral immune response. https://www.news-medical.net/news/20200616/Severity-of-COVID-19-linked-to-heparanase-activity-and-heparan-sulfate-levels.aspx
33/ RaTG13 also presents codon deoptimization. Moreover, its RBM is not able to bind hACE2 and it is not found in any other CoV known. RaTG13 could be therefore the unfinished result of a different vaccine strategy, such as Virus-like particle (VLP).
34/ Development of several vaccine strategies against BtCoV was an active field of research at the WIV and DARPA before SARS2 outbreak. https://twitter.com/BillyBostickson/status/1312049914206056448
35/ Recent ongoing research also involved experiments on self-spreading vaccines. The risk that an unfinished vaccine might have leaked during tests on animal models (e.g. lab worker infection, improper waste disposal) should therefore be not underestimated.
37/ *DRASTIC is a group of independent and not funded researchers working voluntarily together to investigate the origins of SARS2.
https://twitter.com/BillyBostickson/status/1303831074418487296
@CZilcho @ch_zimmer @interne41914499 @schnufi666
https://twitter.com/BillyBostickson/status/1303831074418487296
@CZilcho @ch_zimmer @interne41914499 @schnufi666
38/please unroll @threadreaderapp

 Read on Twitter
Read on Twitter